Why Genomic Signatures Matter
Colorectal cancer (CRC) remains a leading health burden in the United States, with roughly 1.9 million new cases and 900 000 deaths each year. Historically, therapeutic decisions were driven by anatomic stage, tumor size, and nodal involvement. Recent large‑scale whole‑genome and transcriptome studies, however, have revealed distinct molecular subtypes and mutational signatures that supersede anatomy in prognostic and predictive power. Comprehensive profiling of driver mutations (e.g., KRAS, BRAF, RNF43), microsatellite instability, tumor mutational burden, and gene‑expression subtypes such as the five CRPS groups or Consensus Molecular Subtypes enables clinicians to stratify patients into low‑ and high‑risk categories, select anti‑EGFR, BRAF‑targeted, or immunotherapy agents, and even consider de‑intensified adjuvant therapy. Integrating these genomic signatures into treatment algorithms thus personalizes care, improves outcomes, and reduces unnecessary toxicity. Evidence from the 1,063‑tumor genome project and the 981‑sample colibactin study shows that signatures such as SBS44, SBS88, and CRPS1 predict survival and response to inhibitors, underscoring the relevance of molecular classification in colorectal cancer for therapeutic decision‑making.
Genomic Foundations of Colorectal Cancer

Whole‑genome sequencing discoveries
Large‑scale WGS of >1,000 CRCs revealed 96 significantly mutated driver genes, 24 previously unlinked to cancer, and two co‑mutation patterns in the RTK‑WNT axis: KRAS‑APC‑AMER1 and BRAF‑RNF43.
Mutational signatures and clinical relevance
Signature SBS‑CRC1 (MSI/POLE‑associated) and COSMIC SBS44 (hypermutated, right‑sided, BRAF‑V600E) predict longer overall survival. Bacterial colibactin signatures SBS88 and ID18 are enriched in early‑onset CRC, especially before age 40, and drive APC indels, suggesting a microbial contribution to early driver events.
Structural variants, CNVs, and mtDNA mutations
Deletions dominate SVs, hitting tumor‑suppressors (RUNX1, PTEN, and SMAD3). ecDNA amplifications (ERBB2, MYC appear in 24 % of tumours. Mitochondrial truncating mutations (CYB, ND5 occur in ~6 % and correlate with longer survival in non‑hypermutated cases.
Molecular pathways of colorectal cancer
Wnt/β‑catenin dysregulation (APC loss, CTNNB1), MAPK activation (KRAS/NRAS/BRAF), PI3K/AKT (PIK3CA/PTEN), and loss of TP53/SMAD4 drive tumorigenesis, while MSI and CIN create genomic instability.
Signaling pathways: pathogenesis and targeted therapy
Anti‑EGFR antibodies are used in RAS‑wild‑type disease; BRAF‑V600E tumours benefit from combined BRAF/MEK/EGFR inhibition; HER2‑amplified cancers respond to trastuzumab‑based regimens; TGF‑β and stromal activation in CMS4 suggest anti‑angiogenic or stromal‑targeted strategies.
Consensus Molecular Subtypes and Prognostic Gene‑Expression Signatures

Prognostic genome and transcriptome signatures in colorectal cancers
Large‑scale whole‑genome and transcriptome sequencing of >1,000 CRCs has identified driver mutations (WNT, EGFR–KRAS–BRAF, TGFβ, CYB) and copy‑number changes that independently predict overall survival. The COSMIC SBS44 mutational signature, enriched in hypermutated, right‑sided, BRAF‑V600E tumours, and SBS15 (DNA‑repair‑failure) correlate with poorer outcomes. Truncating mtDNA mutations in CYB and ND5 also associate with survival, especially in non‑hypermutated cases.
Molecular pathways of colorectal cancer
Colorectal carcinogenesis follows stepwise alteration of core pathways: APC loss or CTNNB1 activation drives Wnt/β‑catenin signaling; KRAS/NRAS or BRAF mutations hyperactivate MAPK/ERK; PI3K/AKT is potentiated by PIK3CA/PTEN changes. Tumor‑suppressor loss (TP53, SMAD4, MMR genes) yields chromosomal instability or microsatellite instability, often alongside CIMP. These networks underlie the disease’s heterogeneity and guide targeted or immunotherapeutic strategies.
CMS1–CMS4 classification and clinical implications
CMS1 (MSI‑immune) tumours respond to checkpoint inhibitors; CMS2 (canonical) benefit from anti‑EGFR agents when RAS/BRAF wild‑type; CMS3 (metabolic) shows KRAS‑driven reprogramming; CMS4 (mesenchymal) is stromal‑rich, hypoxic, and linked to resistance, indicating need for anti‑angiogenic or stromal‑targeted therapy.
CRPS1–CRPS5 refined subtypes and hypoxia signatures
Gene‑expression profiling defined five prognostic CRC subtypes (CRPS1‑CRPS5) that refine CMS. CRPS1 (hypermutated, MSI, high hypoxia) has the worst post‑recurrence survival; CRPS2/3 enjoy the longest OS/RFS; CRPS4 is stromal‑activated with poor prognosis; CRPS5 shows WNT repression and intermediate outcomes.
Oncotype DX, ColoPrint, and the 12‑gene recurrence score
Validated expression panels (Oncotype DX, ColoPrint, a 12‑gene RNA‑seq recurrence score) stratify stage II/III patients into low‑ and high‑risk groups, informing de‑intensified versus intensified adjuvant chemotherapy decisions.
Prognostic value of mutational signatures and mtDNA mutations
Beyond SBS44/SBS15, novel signatures (SBS‑CRC1, SBS‑CRC2) and regulatory element mutations (ID2, HS3ST1, DAPK1 impact OS/RFS. mtDNA truncating mutations and copy‑number alterations further refine risk, supporting genomic‑signature‑guided treatment algorithms.
Stage‑Specific Treatment Algorithms Guided by Genomic Testing
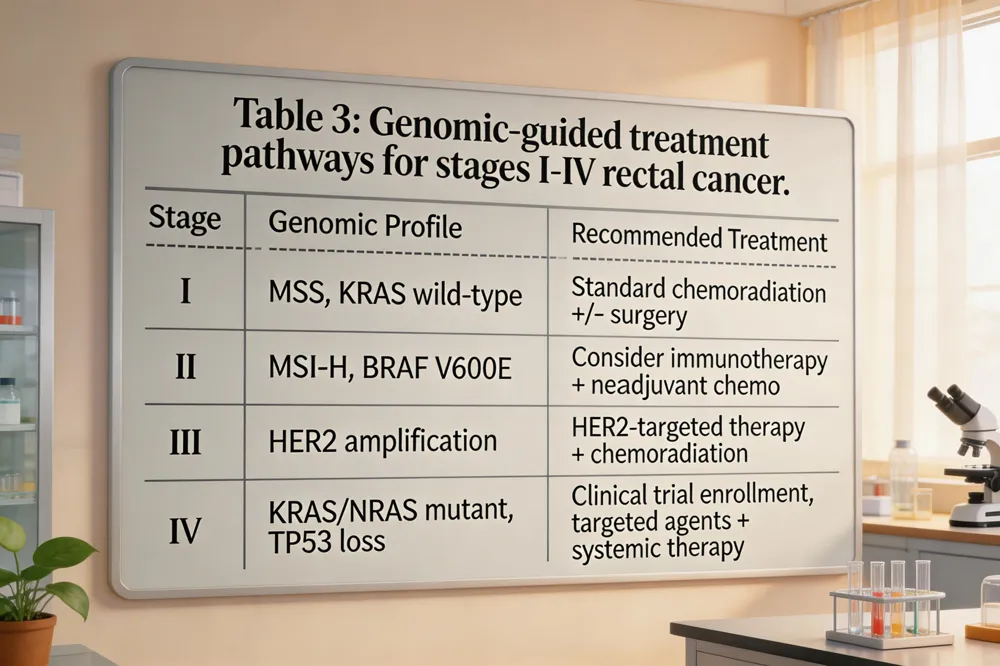

Stage 1 rectal cancer treatment – T1‑T2 N0 M0 lesions are curatively resected. Low‑risk tumors may be removed by transanal local excision (TEM/TAMIS); deeper lesions require total mesorectal excision (TME) with lymph‑node harvest. Adjuvant therapy is reserved for high‑risk pathologic features.
Stage 2 rectal cancer treatment – Multimodal approach: neoadjuvant 5‑FU‑based chemoradiation (or total neoadjuvant therapy, TNT) to downstage, then TME. High‑risk molecular markers (MSI‑H, BRAF‑V600E, KRAS/NRAS mutations, high TMB) guide intensified adjuvant chemotherapy (FOLFOX/CAPEOX) or trial enrollment.
Rectal cancer treatment by stage – Stage 0: local excision only. Stage I: surgery ± adjuvant if adverse pathology. Stages II/III: neoadjuvant chemoradiation + surgery + adjuvant chemo. Stage IV: systemic therapy prioritized; surgery/radiation for palliation or resection of oligometastatic disease.
Rectal cancer treatment stage 4 – Combination chemotherapy (FOLFOX/FOLFIRI) with biologics (anti‑VEGF, anti‑EGFR) and, when MSI‑H or high TMB, immune checkpoint inhibitors (pembrolizumab). Genomic profiling (KRAS/NRAS, BRAF, HER2, NTRK, TMB) directs targeted agents; ctDNA guides adjuvant escalation. Multidisciplinary care ensures therapy aligns with tumor biology and patient goals.
Precision Oncology in Practice: Testing, Biomarkers, and Impact

Precision oncology meaning Precision oncology tailors prevention, diagnosis, and treatment to a patient’s unique tumor DNA, RNA, and protein alterations, enabling selection of targeted therapies, immunotherapies, or dose‑adjusted regimens.
Precision Oncology impact Factor The journal Precision Oncology reported a 2023 Impact Factor of ~4.5, reflecting solid citation performance in personalized cancer medicine.
What is a molecular test for colorectal cancer? Core tests include MSI testing (detecting DNA‑repair defects) and targeted NGS panels for KRAS, NRAS, BRAF, HER2, and NTRK alterations, guiding immunotherapy eligibility and anti‑EGFR use.
Why don't doctors like Cologuard? Its lower detection rate for large polyps (~42 %) and need for confirmatory colonoscopy limit confidence compared with colonoscopy’s ~95 % detection.
ASCO rectal cancer Guidelines 2024 ASCO recommends MRI staging, total neoadjuvant therapy for MSS/T3‑T4 disease, immunotherapy for MSI‑H/dMMR tumors, and watch‑and‑wait for clinical complete responders.
Molecular tests & utility TMB >10 mut/Mb and ctDNA MRD testing identify patients for checkpoint blockade or adjuvant escalation.
Guideline recommendations NCCN 2024 mandates universal MSI, KRAS/NRAS, BRAF, HER2, and NTRK testing; NGS panels are standard for stage II‑III disease.
Gene‑expression assays & cost‑effectiveness Oncotype DX and ColoPrint stratify recurrence risk, allowing de‑intensified chemotherapy in low‑risk patients and reducing toxicities and costs.
Microbial Mutational Signatures and Early‑Onset Disease

Colibactin, a genotoxic metabolite produced by pks⁺ Escherichia coli creates a distinctive DNA‑damage pattern in colorectal tumors. The dominant single‑base‑substitution feature, SBS88 shows T > G transversions in a TpC context, while the accompanying indel signature, ID18 consists of short deletions at homopolymer runs. Both signatures are markedly enriched in early‑onset colorectal cancers (CRC); SBS88 and ID18 appear 2.5‑ to 4‑fold more frequently in patients diagnosed before age 40, a three‑fold enrichment compared with cases after age 70. Whole‑genome sequencing of 981 CRCs from 11 countries revealed that regions with higher age‑standardized incidence — such as parts of Europe and North America — show a greater prevalence of these colibactin‑related signatures, implicating geographic differences in gut‑microbiome exposure. The signatures frequently co‑occur with APC driver mutations, suggesting that colibactin exposure in childhood or early adulthood initiates tumorigenesis. Recognizing this microbial contribution opens avenues for prevention (e.g., microbiome modulation, targeted antibiotic stewardship) and surveillance strategies that focus on high‑risk, young populations in high‑incidence locales.
Future Directions: Integrated Genomics, ctDNA, and Adaptive Trials

Circulating tumor DNA (ctDNA) analysis after resection has emerged as a sensitive minimal‑residual‑disease (MRD) assay; a positive ctDNA result predicts early recurrence and can trigger adjuvant therapy escalation, while a negative result may spare patients unnecessary treatment. Adaptive neoadjuvant strategies such as total neoadjuvant therapy (TNT) and watch‑and‑wait are increasingly guided by molecular response, allowing clinicians to modulate chemoradiation intensity or omit surgery when ctDNA and imaging show deep remission. The co‑evolution of the genome and epigenome in colorectal cancer drives heterogeneity: mutations in chromatin‑modifier genes and recurrent alterations in regulatory DNA access cooperate with DNA‑damage signatures, reshaping transcription‑factor networks (e.g., SOX, HOX, loss of CTCF) and influencing therapeutic resistance. Recognizing hereditary risk, the “3/2:1 rule” – three affected relatives, spanning two generations, with at least one diagnosis before age 50 – remains a practical clinical criterion for Lynch‑syndrome screening and informs cascade testing. Integrating these genomic and epigenomic insights promises more precise, dynamic treatment algorithms.
Putting Genomic Signatures to Work at Hirschfeld Oncology
At Hirschfeld Oncology, multidisciplinary tumor boards convene weekly to review each case with an integrated molecular dossier that includes whole‑genome sequencing, RNA‑seq transcriptomics, and circulating‑tumor‑DNA (ctDNA) profiling. The board’s oncologists, pathologists, radiologists, and bioinformaticians translate mutational signatures—such as MSI‑H, KRAS/NRAS wild‑type, BRAF V600E, CMS subtypes, and high tumor‑mutational burden—into concrete therapeutic recommendations. By matching these signatures to FDA‑approved targeted agents or immunotherapies, the team can safely de‑intensify adjuvant chemotherapy for low‑risk patients, sparing them unnecessary toxicity, while escalating therapy (e.g., combined BRAF/MEK/EGFR inhibition, anti‑angiogenic regimens, or checkpoint blockade) for high‑risk genomic profiles. Hirschfeld Oncology also enrolls eligible patients in genotype‑driven clinical trials and continuously updates its treatment algorithms based on emerging data, ensuring that each patient receives a personalized, evidence‑based plan that maximizes efficacy and minimizes overtreatment.





.png)


.png)
.png)